1)The first major step for the DNA Replication to take place is the breaking of hydrogen bonds between bases of the two antiparallel strands. The unwounding of the two strands is the starting point. The splitting happens in places of the chains which are rich in A-T. That is because there are only two bonds between Adenine and Thymine (there are three hydrogen bonds between Cytosine and Guanine). Helicase is the enzyme that splits the two strands. The initiation point where the splitting starts is called "origin of replication".The structure that is created is known as "Replication Fork"
.
2) One of the most important steps of DNA Replication is the binding of RNA Primase in the the initiation point of the 3'-5' parent chain. RNA Primase can attract RNA nucleotides which bind to the DNA nucleotides of the 3'-5' strand due to the hydrogen bonds between the bases. RNA nucleotides are the primers (starters) for the binding of DNA nucleotides.
3) The elongation process is different for the 5'-3' and 3'-5' template. a)5'-3' Template: The 3'-5' proceeding daughter strand -that uses a 5'-3' template- is called leading strand because DNA Polymerase ä can "read" the template and continuously adds nucleotides (complementary to the nucleotides of the template, for example Adenine opposite to Thymine etc).
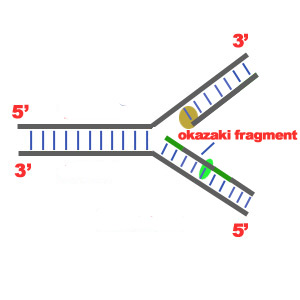
b)3'-5'Template: The 3'-5' template cannot be "read" by DNA Polymerase ä. The replication of this template is complicated and the new strand is called lagging strand. In the lagging strand the RNA Primase adds more RNA Primers. DNA polymerase å reads the template and lengthens the bursts. The gap between two RNA primers is called "Okazaki Fragments".
The RNA Primers are necessary for DNA Polymerase å to bind Nucleotides to the 3' end of them. The daughter strand is elongated with the binding of more DNA nucleotides.

I appreciate all of the information that you have shared. Thank you for the hard work!
ReplyDeleteKlenow (3′→ 5′ exo-) is a mesophilic dna polymerase deficient in both proofreading (3′→ 5′) and nick-translation (5′→ 3′) nuclease activities, and that displays a moderate strand displacement activity during DNA synthesis.